|
|
|
Copyright © 2002 by the author(s). Published here under license by The Resilience Alliance.
Go to the pdf version of this article.
The following is the established format for referencing this article:
Spong, G. and L. Hellborg. 2002. A near-extinction event in lynx: do microsatellite data tell the tale? Conservation Ecology 6(1): 15. [online] URL: http://www.consecol.org/vol6/iss1/art15/
A version of this article in which text, figures, tables, and appendices are separate files may be found by following this link.
Insight A Near-extinction Event in Lynx: Do Microsatellite Data Tell the Tale? Goran Spong and Linda Hellborg
Evolutionary Biology Centre, Uppsala University
- Abstract
- Introduction
- Methods and Materials
- Results and Discussion
- Responses to this Article
- Acknowledgments
- Literature Cited
Fluctuations in population size can have a profound impact on effective population size and the maintenance of genetic variation within a population. A number of tests based on microsatellite data have been developed for the detection of bottleneck events in a population's past. In this paper, we search for signs of a bottleneck in microsatellite data on the Scandinavian lynx (Lynx lynx) population. This population was hunted to the brink of extinction, with fewer than 100 animals (one estimate was as low as 30 individuals) remaining in the late 1920s. Protection allowed recovery of the population, which currently numbers about 2000 adults. Despite this pronounced demographic bottleneck (more than 95% of the population was killed), we could find no conclusive genetic evidence of a bottleneck using four of the recently developed tests for detecting such events.
KEY WORDS: bottleneck, conservation, extinction, genetic testing, lynx, microsatellite data, population history, Scandinavia.
Published: June 27, 2002
A number of statistical methods now make it possible to investigate a population's history without the need for information on past population sizes (e.g., Kimmel et al. 1998; Luikart et al. 1998a, 1998b; Reich et al. 1999; Spencer et al. 2000). These tests typically quantify deviations from expected patterns in allele sizes, allele numbers, heterozygosity levels, or allele distributions, often using microsatellite data. These methods hold promise because information about a population's past history is often useful when predicting future population dynamics and extinction risks resulting from demographic or genetic processes.
Two commonly cited examples of bottlenecked populations are the lions (Panthera leo) of the Ngorongoro crater in Tanzania (see Packer et al. 1991) and the northern elephant seals (Mirounga augustirostrus) on the west coast of North America (see Hedrick 1995, Hoelzel 1999). In Ngorongoro, only seven females and eight males founded the current population. Consequently, these crater lions have significantly lower genetic variation than the large population in nearby Serengeti, and the males exhibit a high proportion of abnormal sperm, presumably due to inbreeding effects (Packer et al. 1991). North American elephant seals are thought to have gone through a bottleneck of 10–20 individuals and also show low genetic variation (Hedrick 1995, Hoelzel 1999). Both populations have recovered and are currently protected, but their low genetic variation may, for example, result in inbreeding depression (e.g., Barone et al. 1994, Frankham 1995) or make them more sensitive to pathogens (e.g., O'Brien 1994). The crater lion population reached 124 individuals in the early 1980s, but has been in constant decline since then and now numbers only 54 lions (with a mere 11 adults). The cause or causes of the decline are not known, but disease is the alleged culprit in several cases (C. Packer, personal communication).
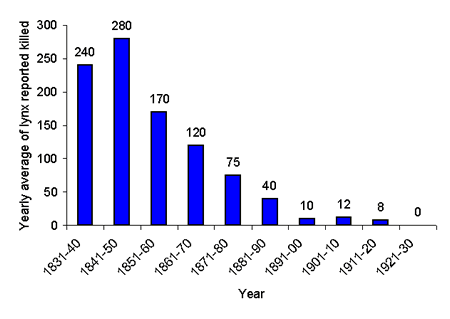
Lynx (Lynx lynx) are medium-sized, solitary felids with a holarctic distribution. Lynx recolonized the Scandinavian Peninsula after the last ice age and were probably abundant until the beginning of the 19th century, when a significant population decline began after the introduction of efficient firearms (see Fig. 1). Extensive winter surveys showed that the total number of animals in Scandinavia at the end of the 1920s was fewer than 100 (adults and dependent offspring), and a reasonable estimate indicated about 30 individuals (Lönnberg 1930). Despite the great economic value of lynx pelts, no lynx were trapped or shot during the 1920s; this suggests that the population at the time was very small and supports the lower estimate. Today the population numbers about 2000 and has most likely been isolated since the recovery, with no gene flow from the neighboring Finnish population (see the Results and Discussion sections and Hellborg et al. 2002). The Scandinavian lynx population thus presents a good opportunity to test for signs of a bottleneck in genetic data.
Tissue samples from 89 Swedish and 107 Norwegian lynx were collected from animals legally shot between the years 1993 and 1998. These samples consisted of muscle or tissues from internal organs stored in ethanol or frozen at -20ºC. Extractions followed a standard phenol/chloroform protocol (Sambrook et al. 1989).
Eleven unlinked polymorphic microsatellite loci and one monomorphic locus were scored (Hellborg et al. 2002) using primers developed for the domestic cat Felis catus (Menotti-Raymond et al. 1999). Typical polymerase chain reaction (PCR) testing was carried out under these conditions: 25 ng of target DNA was used in 10-µl PCR reactions containing 1 x PCR buffer II (Perkin Elmer), 5 µM of each primer, 16 mM dNTP, and 0.5 U AmpliTaq polymerase (Perkin Elmer). The PCR profile included one cycle at 94ºC for 3 min, followed by 32 cycles at 94ºC for 30 s, 55ºC for 30 s, and 72ºC for 1 min, ending at 72ºC for 10 min. Alleles were scored using either fluorescently labeled primers with products analyzed on an ABI377 (Perkin Elmer) instrument, radioactive labeled primers with products visualized by autoradiography (Sambrook et al. 1989), or unlabeled primers with products visualized by silver staining (Bassam et al. 1991). Some samples were analyzed by all three methods to ensure consistency.
Statistical methodsThe bottleneck tests were performed using BOTTLENECK software (Cornuet and Luikart 1996) and two tests described in Reich et al. (1999). The first test in BOTTLENECK is based on the fact that reductions in effective population size result in losses of alleles and heterozygosity. Because allele numbers are reduced more quickly than gene diversity, a bottleneck can be detected by comparing the two.
The second method in BOTTLENECK plots the allele frequency distribution and compares it with the expected L-shaped distribution. A population that has been through a bottleneck is likely to exhibit a mode shift in the frequency distribution of alleles.
The tests introduced by Reich et al. (1999), the intralocus variance k test and the interlocus g test, differ from the first two tests in that they detect population expansion rather than the actual bottleneck. Both tests assume that microsatellite loci evolve according to the stepwise mutation model, although the g test can be modified to apply to any genetic marker. The k test is based on the assumption that a constant-sized population will have several modes in its allele length distribution (due to multiple ancient branching events in the gene genealogy at each locus), whereas an expanding population is expected to have a single mode and be more peaked.
The g test takes advantage of the observation that the variance in the widths of allele length distribution across loci is usually lower in an expanding population compared to a population that remains constant in size. The derived variance of the variances across loci can thus be compared to what is theoretically expected.
All loci were in Hardy Weinberg (HW) equilibrium as measured in GENEPOP (Raymond and Rousset 1995) and showed low heterozygosity levels with an average of 0.49 ± 0.06 (mean ±SE henceforth) and an average number of alleles of 4.7 ± 0.7 (Table 1). This can be compared to heterozygosity levels of 0.62 for lynx in Finland (Hellborg et al. 2002), 0.60 for lynx from the Baltic states (Hellborg et al. 2002), 0.66 for lynx in North America (Schwartz et al. 2002), 0.77 for the leopard Panthera pardus (Spong et al. 2000), and 0.69–0.75 for lions in Tanzania (Spong 2001). Despite the fact that we screened samples from 176 Scandinavian lynx but only 80 samples from lynx from Finland and the Baltic states, the mean number of alleles per loci was lower in Scandinavia: 4.7 and 5.3, respectively.
|
Table 1. Statistics of the loci used in the analyses. Hobs stands for "observed levels"; HexpIAM stands for "expected value under the infinite allele model (IAM)," and HexpSMM for "expected value under the stepwise mutation model (SMM)," i.e., the values that would have been expected had the loci evolved under these two models.
† Deviates significantly from values obtained using GENEPOP software (Raymond and Rousset 1995).
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The population showed weak signs of gene flow with the Finnish population, i.e., 0.82 individuals per generation (Hellborg et al. 2002). However, estimates of gene flow cannot differentiate between past and present gene flow. Observational data strongly suggest a complete absence of lynx in northern Scandinavia until recently. Likewise, lynx have been absent from northern Finland for decades and still are. Thus, it seems reasonable to assume that the small Nm value derived is explained by historic dispersal events that took place before 1920, i.e., prebottleneck.
Depending on the assumed mutation model, a few loci showed significant deviations from expected heterozygosity levels (see Table 1). Two loci showed significant excess under the infinite allele model, while one locus showed significant deficit under the stepwise mutation model. Similarly, the heterozygosity excess method implemented by BOTTLENECK produced highly different results depending on the mutation model. Under the assumption that the loci evolved according to an infinite allele model, the test detected a bottleneck (P = 0.006), whereas, under a stepwise mutation model, there was no support for a bottleneck (P = 0.12). The intermediate model, called the two-phased mutation model, did detect a bottleneck if the proportion of single stepwise mutations was below about 60%, with the remainder occurring as multistep (with a variance that concurred with the variance in the data set used). Because microsatellite loci have been shown to conform better to the stepwise mutation model (Shriver et al. 1993) than to the infinite allele model, the detection method based on the infinite allele model was considered to be invalid. Although estimates of the proportion of multistep mutations for microsatellite loci vary, the proportion is unlikely to significantly exceed about 10% (Ellegren 2000). Having to set the proportion of multistep mutations to > 40% before significance is achieved thus also invalidated this result.
The second test in BOTTLENECK did not detect any mode shift in the frequency distribution of alleles. Our data set was found to conform to what is expected in a population that has not experienced a bottleneck (normal L-shaped curve), and this test consequently showed no signs of a bottleneck.
The k test did not detect the recent expansion, because only four of 12 loci had a positive kurtosis, which is not significantly different from the expected binomial distribution (P = 0.22).
The g test exhibited a value of 0.90. For this test to indicate a bottleneck, the value has to be below 0.24 at P < 0.05 (P = 160) as tabulated by Reich et al. (1999).
Despite convincing observational evidence of a demographic bottleneck and subsequent recovery of the Scandinavian lynx population, all the genetic tests we applied failed to detect any signs of this bottleneck in our microsatellite data. The persecution of the Scandinavian lynx reduced the population by more than 95%, perhaps even 99% (assuming ~3000 lynx preceding a bottleneck of 30 individuals). Even the conservative estimate of 95% is a very narrow bottleneck, and of a size that could have profound implications for any large mammal, e.g., disrupting social groups, changing distribution patterns, or altering population age structure or sex ratio.
What might have caused this detection failure? First, the population size during the bottleneck may have been underestimated, and thus the bottleneck not as severe as alleged. However, because of the great economic value put on lynx pelts in the past, we feel that hunting statistics present convincing evidence of the severity of the decline, and we therefore find this explanation unlikely. Second, there may have been undetected migration in recent times (i.e., during the latter half of the 20th century) from the Finnish lynx population, which would have diminished the effects of the bottleneck. However, genetic analyses of Finnish lynx show a very low estimate of gene flow between Sweden and Finland, and we believe that this number reflects ancient migration events that took place before the bottleneck. The last and most probable explanation concerns the number of loci used. The use of only a few loci restricts the power of the tests we chose, although our 11 polymorphic loci exceeded the minimum of 10 recommended for the BOTTLENECK tests. To achieve powers of 0.5 or more for the k test and the g test, more than 30 loci are needed (Reich et al. 1999). Our data set of 12 loci resulted in a power of between 0.1 and 0.4. Few studies have scored the number of loci required to attain reasonable power, presumably because reliable relatedness estimates and/or F-statistics can be obtained with far fewer loci (and analyses are costly). However, keep in mind that a minimum of 10 polymorphic loci is recommended for the heterozygote excess method implemented in BOTTLENECK. In a performance test of this method on 11 microsatellite data sets from populations known to have experienced a demographic bottleneck and scored at four to 10 loci, signs of a bottleneck were detected in nine of these under the infinite allele model. Assuming a stepwise mutation model, only five of the populations showed signs of a bottleneck (Luikart and Cornuet 1998). All but one of these populations were scored for fewer than 10 loci, and the population scored at 10 loci showed no signs of a bottleneck. This is similar to our study findings, where, despite having 11 polymorphic loci, we failed to detect a bottleneck.
In conclusion, bottlenecks that are short or moderately narrow may have little impact on patterns of genetic variation. This may render the bottleneck invisible to the tests described above, even when the desired number of loci has been included. Because of this, caution is advisable. Bottlenecks of severe nature from a demographic viewpoint may often evade detection. Stochastic events during times of small population size are probably more of a threat than the depletion of genetic variation for most populations entering bottlenecks. Our data had power comparable to data sets available for many other species, so a lack of power in bottleneck tests using microsatellite data is likely to be a widespread problem.
Responses to this article are invited. If accepted for publication, your response will be hyperlinked to the article. To submit a comment, follow this link. To read comments already accepted, follow this link.
We thank Mats Björklund, Scott Creel, and Carles Vila for help and constructive comments on the manuscript. GS is supported by a Ph.D. grant from the Swedish International Developmental Agency.
Barone, M. A., M. E. Roelke, J. Howard, J. L. Brown, A. E. Anderson, and D. E. Wildt. 1994. Reproductive characteristics of male Florida panthers: comparative studies from Florida, Texas, Colorado, Latin America, and North American zoos. Journal of Mammalogy 75:150-162.
Bassam, B. J., A. G. Caetano, and P. M. Gresshoff. 1991. Fast and sensitive silver staining of DNA in polyacrylamide gels. Analytical Biochemistry 196:80-83.
Cornuet, J. M., and G. Luikart. 1996. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001-2014.
Ellegren, H. 2000. Heterogeneous mutation processes in microsatellite DNA sequences. Nature Genetics 24:400-402.
Frankham, R. 1995. Inbreeding and extinction: a threshold effect. Conservation Biology 9:792-799.
Hedrick, P. W. 1995. Elephant seals and the estimation of a population bottleneck. Journal of Heredity 86:232-235.
Hellborg, L., C. W. Walker, E. K. Rueness, J. E. Stacy, I. Kojola, H. Valdmann, C. Vila, B. Zimmermann, K. S. Jakobsen, and H. Ellegren. 2002. Differentiation and levels of genetic variation in northern European lynx (Lynx lynx) populations as revealed by microsatellite and mitochondrial DNA analysis. Conservation Genetics 3:97-111.
Hoelzel, A. R. 1999. Impact of population bottlenecks on genetic variation and the importance of life-history; a case study of the northern elephant seal. Biological Journal of the Linnean Society 68:23-39.
Kimmel, M., R. Chakraborty, J. P. King, M. Bamshad, W. S. Watkins, and L. B. Jorde. 1998. Signatures of population expansion in microsatellite repeat data. Genetics 148:1921-1930.
Liberg, O. 1997. Lodjuret. Pages 9-20 in O. Liberg. Historik och utbredning. Almqvist and Wiksell, Uppsala, Sweden.
Lönnberg, E. 1930. Lodjurets förekomst i Sverige de sista hundra åren. Kungliga Vetenskapsakademiens skrifter i naturskyddsärenden, Number 14. Kungliga Vetenskapsakdemiens, Stockholm, Sweden.
Luikart, G. L., and J-M. Cornuet. 1998. Empirical evaluation of a test for identifying recently bottlenecked populations from allele frequency data. Conservation Biology 12: 228-237.
Luikart, G. L., W. B. Sherwin, B. M. Steele, and F. W. Allendorf. 1998a. Usefulness of molecular markers for detecting population bottlenecks via monitoring genetic change. Molecular Ecology 7:963-974.
Luikart, G. L., W. B. Sherwin, B. M. Steele, and F. W. Allendorf. 1998b. Distortion of allele frequency distributions provides a test for recent population bottlenecks. Journal of Heredity 89:238-247.
Menotti-Raymond, M., V. A. David, L. A. Lyons, A. A., Schaffer, J. F. Tomlin, M. K. Hutton, and S. J. O' Brien. 1999. A genetic linkage map of microsatellites in the domestic cat (Felis catus). Genomics 57:9-23.
O'Brien, S. J. 1994. A role for molecular genetics in biological conservation. Proceedings of the National Academy of Sciences 91:5748-5755.
Packer, C., A. E. Pusey, H. Rowley, D. A. Gilbert, J. Martenson, and S. J. O'Brien. 1991. Case study of a population bottleneck: lions of the Ngorongoro crater (Tanzania). Conservation Biology 5:219-230
Raymond, M., and F. Rousset. 1995. GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Journal of Heredity 86:248-249.
Reich, D. R., M. W. Feldman, and D. B. Goldstein. 1999. Statistical properties of two tests that use multilocus data sets to detect population expansions. Molecular Biology and Evolution 16:453-466.
Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual. Second edition. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA.
Schwartz, M. K., L. S. Mills, K. S. McKelvey, L. F. Ruggiero, and F. W. Allendorf. 2002. DNA reveals high dispersal synchronizing the population dynamics of Canada lynx. Nature 415:520-522.
Shriver, M. D., L. Jin, R. Chakraborty, and E. Boerwinkle. 1993. VNTR allele frequency distributions under the stepwise mutation model: a computer simulation approach. Genetics 134:983-993.
Spencer, C. C., J. E. Neigel, and P. L. Leberg. 2000. Experimental evaluation of the usefulness of microsatellite DNA for detecting demographic bottlenecks. Molecular Ecology 9:1517-1528.
Spong, G. 2001. Genetic consequences of dispersal and social behavior in lions, Panthera leo. Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology, Number 659. Uppsala University, Uppsala, Sweden.
Spong, G., M. Johansson, and M. Björklund. 2000. High genetic variation in leopards indicates large and long-term stable effective population size. Molecular Ecology 9:1773-1782.
Address of Correspondent:
Goran Spong
Evolutionary Biology Centre
Uppsala University
Norbyvägen 18D
Uppsala SE-752 36, Sweden
Homepage of Goran Spong
Phone: +46 18 471 26 34
Fax: +46 18 471 64 84
goran.spong@ebc.uu.se