|
|
|
Copyright ©1998 by The Resilience Alliance*
Francisco Rodríguez-Trelles, Miguel A. Rodríguez and Samuel M. Scheiner. 1998. Tracking the genetic effects of global warming: Drosophila and other model systems. Conservation Ecology [online] 2(2): 2. Available from the Internet. URL: http://www.consecol.org/vol2/iss2/art2/
A version of this article in which text, figures, tables, and appendices are separate files may be found by following this link.
Insight Tracking the genetic effects of global warming: Drosophila and other model systems Francisco Rodríguez-Trelles1, Miguel A. Rodríguez2, and Samuel M. Scheiner3
1Departament de Genètica i de Microbiología, Universitat Autonoma de Barcelona, Spain; 2Area de Ecologia, Facultad de Ciencias, Universidad de Alcala, Spain; 3Arizona State University West, USA
- Abstract
- General Issues
- Microevolution in D. subobscura
- The Genus Drosophila as a Model System
- Other Species
- Conclusions
- Responses to this Article
- Acknowledgments
- Literature Cited
Current efforts to study the biological effects of global change have focused on ecological responses, particularly shifts in species ranges. Mostly ignored are microevolutionary changes. Genetic changes may be at least as important as ecological ones in determining species' responses. In addition, such changes may be a sensitive indicator of global changes that will provide different information than that provided by range shifts. We discuss potential candidate systems to use in such monitoring programs. Studies of Drosophila subobscura suggest that its chromosomal inversion polymorphisms are responding to global warming. Drosophila inversion polymorphisms can be useful indicators of the effects of climate change on populations and ecosystems. Other species also hold the potential to become important indicators of global change. Such studies might significantly influence ecosystem conservation policies and research priorities.
KEY WORDS: biodiversity conservation, biological indicator, chromosomal inversion polymorphism, climate change,
The current greenhouse-induced increase in world temperatures is exceedingly rapid compared with prehistoric climate changes of similar magnitude (Vitousek 1992, Schneider 1993). In the face of such global environmental change, adaptive evolution may be the only route to many species' long-term survival (Travis and Futuyma 1993, Lande and Shannon 1996). In spite of their potential relevance, microevolutionary responses have, however, been largely neglected in detecting biological consequences of global warming. Most studies assume that worldwide climate amelioration will lead primarily to shifts in species' geographic boundaries (Schneider 1993), and have correspondingly focused on tracking population movements at species borders (Parmesan 1997), with little or no interest in genetic analyses. Although some already claim that climate warming is affecting species' distributions (Parmesan 1997), we know little about microevolutionary responses, which may occur instead of, or in conjunction with, boundary shifts. Because species differ in their ability and rate of migration (Davis 1981), both demographic and evolutionary processes need to be studied.
At least some species have the potential to experience substantial adaptive responses to climate warming. In turn, these rapid adaptive responses might affect genetic or chromosomal diversity at some localities. Such responses are scarcely surprising, given the many examples in plants and animals that document rapid adaptation to environmental change (Endler 1986, Travis and Futuyma 1993). However, because of the recent emphasis on boundary shifts, the potential importance of genetic change seems to have been forgotten.
Identifying sensitive and reliable indicators for assessing levels of environmental change is a priority for population and community research on global change (Kingsolver et al. 1993, Bella et al. 1994). However, solely monitoring species dynamics is insufficient if we are to understand biotic responses to global change. Efforts must take into account variation within and among populations. For example, population dynamics are responsive to changes in gene frequencies (e.g., Richardson 1974, Kingsolver and Watt 1983, Watt et al. 1983, Hilbish and Koehn 1985, Templeton et al. 1990). Thus, the ultimate responses of the world's species to global change will depend on levels of genetic variation.
In this paper, we examine the issue of monitoring microevolutionary change as a way of studying the effects of global change. We look at this issue at several levels. At the most detailed level, we consider recent data on Drosophila subobscura. Changes in chromosomal inversion frequencies in this species present the clearest evidence to date of a microevolutionary response to global change (Rodríguez-Trelles and Rodríguez 1998). This species also holds great promise as a model system for such studies over much of the world. At a more general level, we examine the potential for the genus Drosophila to serve as a broad-scale indicator group. Finally, we suggest other well-studied species that might also be monitored or measured.
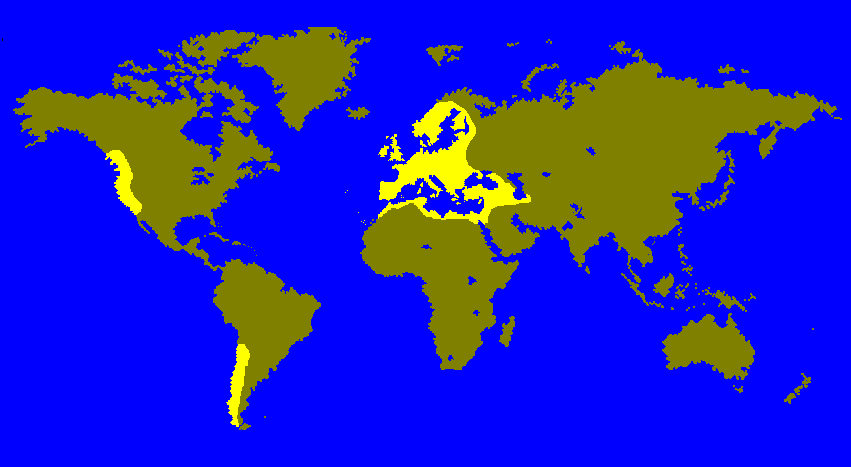
Since the discovery of variation in the polytene chromosomes of Drosophila (Sturtevant and Dobzhansky 1936), inversion polymorphism studies have revealed some of the intricacies of evolutionary change. In most cases, climate-related variables have been invoked as the major factors that determine the frequency of inversions in natural populations. The Palearctic species D. subobscura provides the most conclusive evidence that this is indeed the case (Fig. 1). There are well-established latitudinal clines in the original populations of this species for its five large, acrocentric chromosomes (Krimbas 1992). The clines have also independently evolved in the recently colonized areas of North and South America (Prevosti et al. 1988), strongly suggesting that temperature is the most important variable.
Fig. 1. Worldwide range of Drosophila subobscura. Originally an Old-World species, it has recently colonized North and South America.
|
Extensive research makes Drosophila chromosomal inversion polymorphisms one of the best known genetic systems (Krimbas and Powell 1992). In the particular case of chromosome O of D. subobscura, more than 30 different inversions have been described (Krimbas 1992), many of them showing spatiotemporal heterogeneous distributions (Mennozzi and Krimbas 1992) as well as seasonal cyclic variation (Rodríguez-Trelles et al. 1996). These studies strongly suggest that different inversions carry alleles that are differentially favored in warm and cold climates. Studies of the geographical distribution of its chromosomal polymorphisms in the Palearctic date back to the mid-1950s (Krimbas 1992; see Appendix). About 140 different populations have been investigated throughout this region. From this information, one could build detailed maps of the spatial distribution of the species and its inversions (Krimbas and Loukas 1980, Krimbas 1992, Mennozzi and Krimbas 1992).
Long-term monitoring of inversion frequencies in a Spanish population found seasonal variation that fits expectations based on the latitudinal clines (Rodríguez-Trelles et al. 1996). The seasonal variation is coupled with a long-term decline in the frequency of those inversions presumably carrying alleles associated with adaptation to colder temperatures. We (Rodríguez-Trelles and Rodríguez 1998) observed an 18.3% decrease in inversion diversity of chromosome O during the period 1976 1991. This decline is strongly correlated with the climate-warming that has affected the northern hemisphere since the mid-1970s. Thus, these results conform to projected scenarios on the effect of climate-warming.
We can use this genetic information to better predict the fate of D. subobscura in Iberia and Europe. The key is several inversions that differ in the sensitivity of their alleles to temperature: OST, whose carriers perform better in colder environments, and O3+4, among others, which predominate in warmer environments. Extrapolating from available temporal data in the Iberian peninsula, it appears that the OST inversion, with its alleles that improve performance in the cold, is being replaced there by the inversions that improve warm-temperature performance. This replacement insures the persistence of the species in the Iberian peninsula for now. This adaptive process could not be detected solely by tracking population numbers.
This information also provides insight into species' range changes, as such shifts may be due to microevolutionary changes. In Europe, D. subobscura has shifted toward northern latitudes (Saura 1994). Presumably, shifts in the species' range resulted from flies carrying the OST gene arrangement. Thus, rather than a mere local change in population density or range expansion, the advance of D. subobscura to the north may reflect differences among genotypes in their ability to cope with the new environment. Knowledge of these responses will be critical in understanding the evolutionary potential of species' responses to global warming. Gathering genetic information (e.g., inversion frequencies) along with demographic data may reveal adaptive responses. We emphasize that genetic studies cannot replace demographic and range data, rather that they are a vital complement to them.
Direct, long-term monitoring of some representative populations, with an emphasis on species boundaries, could produce, in the near future, critical information on how climate warming is affecting the spatial distribution of inversions and their rates of change. Drosophila subobscura recently colonized North and South America. The newly occupied territories provide abiotic environments that replicate those in the Palearctic (Prevosti et al. 1988). In addition, several potentially important traits, such as body size and developmental rate, are associated with some gene arrangements in this species (Misra and Reeve 1964, Prevosti 1966). Recording quantitative variation, in conjunction with inversion data, can provide insights on phenotypic responses to climate change. The genetic architecture of responsive quantitative traits could be explored subsequently in the laboratory, using techniques such as QTL mapping. By mapping this variation to different gene arrangements (e.g., OST vs. O3+4, evolutionary changes in important life history traits can be easily tracked.
Additional efforts can focus on relevant major loci. Candidate genes for these analyses are the Hsp70 family. These genes are expressed in response to multiple harmful situations, including abnormally high temperatures. The resulting proteins enhance the probability of survival by increasing resistance to stress. Studies in D. melanogaster showed that differences in the number of Hsp70 gene copies imply dramatic differences in viability to heat shock treatment in the lab (Feder et al. 1996). In D. subobscura, this gene family controls the principal inducible heat-shock protein, and is located on the O3+4 inversion region (89A and 90A loci; Moltó et al. 1992). A challenging hypothesis, yet unexplored, is that observed spatiotemporal patterns of chromosomal polymorphisms reflect differences in the number of heat shock gene copies among inversions. Indeed, differences in copy number have already been found associated with inversions in other gene families (e.g., the amylase region in D. pseudoobscura; Aquadro et al. 1991). Besides the heat shock genes, the clock gene period (per) has recently become a focus for studies of microevolutionary effects of climate warming. In D. melanogaster, the major length variants of the per gene exhibit a clinal distribution in Europe and North Africa that is causally related to the flies' ability to maintain a circadian period at different temperatures (Sawyer et al. 1997). Progress at identifying chromosomal homologies in the Drosophila genome project will no doubt provide extremely valuable information for these studies.
Owing to their relatively narrow physical limits for resource utilization, Drosophila species have been recognized as potentially useful candidates for assessing levels of environmental change (Parsons 1991). Because chromosomal inversions can have large fitness effects (Wright and Dobzhansky 1946, Dobzhansky 1947), they are extremely valuable in monitoring genetic variation for stress-response. In addition, in contrast to single-locus approaches (e.g., allozyme markers), inverted blocks of genes contain multiple loci with linkage and epistatic interactions, thereby providing a more realistic basis for exploring phenotypic responses in potentially key ecological traits.
Comprehensive data on chromosomal inversion polymorphisms exist for a number of Drosophila species, including the closely related nearctic D. pseudoobscura, the nearctic D. robusta, and D. repleta; the neotropical D. willistoni; the cosmopolitan D. melanogaster; and the subcosmopolitan D. buzzatii. Detailed information on some of these species can be found in Krimbas (1992). For exhaustive lists of published literature on Drosophila, see Flybase Home Page, and The Drosophila Virtual Home Page.
On the technical side, use of Drosophila chromosomal polymorphisms could benefit from methods for fly sampling, and chromosomal isolation and identification, fly marker stock availability, techniques for chromosomal staining, and maps of chromosomal inversions, which are easy to carry out and are already optimized and standardized across laboratories. This standardization simplifies comparisons among research groups.
Besides its potential advantages, using chromosomal inversion polymorphisms to monitor the biological consequences of climate change also faces some limitations. The most serious are current limits on our understanding of the ecology of Drosophila and how it is connected to evolutionary processes. Measuring gene flow among populations and the capacity to colonize new habitats are examples of this concern. These aspects can critically affect population structure and the ability of a species to exploit its total genetic variation to respond to novel conditions (Travis and Futuyma 1993). We still need to determine which key ecological factors limit species' distributions, and which phenotypic traits determine survival under rapid environmental change
We have emphasized the genus Drosophila because it is such a well-studied system worldwide. Other groups, however, have much the same potential. We describe a few here to indicate potential research avenues. In general, any such group should have the following properties: (1) a relatively well-studied genome and ecology; (2) easily monitored genetic markers; and (3) a relatively short generation time, so that microevolutionary responses are likely to be observed.
Among insects, grasshoppers are both important ecologically and well-studied genetically. In particular, White's (1978) extensive studies of cytogenetic patterns in Australia might provide a system similar to that we have just described for easy monitoring of genetic changes. A number of small rodents, particularly in the genera Mus and Peromyscus, are well-studied both ecologically and genetically in both Europe and North America (see the Virtual Library of Genetics Home Page, and the Peromyscus Database Home Page). Among plants, the genus Arabidopsis is a prime candidate. Arabidopsis thaliana has one of the best characterized genomes among plant species (see the Arabidopsis Home Page for data on the genetics, ecology, and distribution of Arabidopsis). It has become a cosmopolitan weed, although its extreme southern and northern distribution limits, predicted to be more sensitive to global warming, remain to be determined (Dr. Robert Price, personal communication). Other members of the genus are distributed across the northern hemisphere. Among aquatic species, the genus Daphnia is particularly promising (Peters and De Bernardi 1987). Ecologically, it is a keystone genus. Its genetics has been extensively studied in both North America and Eurasia. This list is not exhaustive. Rather, it indicates the large potential for the approach described here. In some cases, current information is sufficient that population monitoring could begin immediately. In other cases, very little additional information would be needed to make these groups suitable candidates. What is most important is that we recognize the need to monitor genetic changes within species.
In situations in which a species cannot avoid an increasingly stressful environment by shifting its geographic distribution, adaptation or extinction will rely decisively upon its ability to mobilize sufficient genetic variation to track the environmental change (Travis and Futuyma 1993, Lande and Shannon 1996). The potential for adaptive responses to strong, directional selection imposed by climate change may be limited, either because additive genetic variance for the selected traits becomes exhausted, or because it is counterbalanced by genetically correlated responses in other characters (Holt 1990, Travis and Futuyma 1993). Accordingly, taking into account that the probability of fixation of deleterious pleiotropic alleles declines as population size increases (Crow and Kimura 1970), larger populations are expected to have greater responses to climate warming than do smaller populations, which are prone to accumulate a greater mutational load (Holt 1990, Travis and Futuyma 1993, Lynch et al. 1995). This implies that intensities of directional selection capable of inducing microevolutionary responses in a generalist species like D. subobscura, with large population numbers, wide geographic distribution, and high dispersal rates (Krimbas 1992), may ultimately drive to extinction many rarer, specialist, and noncolonist species. Therefore, conservation strategies and efforts by taxonomists to record extant biodiversity in the face of the ongoing climate warming, should be urgently directed to these species and their potential habitats.
The approach outlined here could further provide a potentially fertile experimental link between microevolutionary responses and the dynamics of ecosystems (Holt 1995). Insects are important contributors to nutrient and carbon cycling, to energy flow, and to biomass decomposition (Schowalter et al. 1986). Consequently, changes in insect species' genetic composition induced by climate warming may have profound consequences on ecosystem processes through their influence on population growth rates, phenology, and colonization ability. Experimental results in D. subobscura showed that flies carrying gene arrangements that prevail in southern populations exhibited higher developmental rates than those carrying gene arrangements that are common in northern populations (Krimbas 1992). Genetic differences for this and other life history traits are doubtless being filtered by current levels of climate warming in other insect species as well. For instance, changes like these already may be altering threshold disturbance regimes, perhaps as outbreaks of insect pests.
Species do not exist in isolation, but in dynamic relationships with competitors, predators, pathogens, and parasites; these interactions, particularly competition, are all sensitive to temperature (Jenkinson et al. 1996, Davis et al. 1998). Studies of the drosophilid fauna in Chile show that, coinciding with the recent expansion of D. subobscura into the area, the relative abundance of other species (e.g., D. pavani and D. inmigrans) has dramatically declined, presumably outcompeted by the new colonist (Brncic 1994). Similarly, the presumptive climate-mediated displacement of D. subobscura northward in Europe is expected to alter the structure of local communities.
Evolutionary biologists can provide a unique perspective to the examination of how climate change will affect the earth's biota. It is increasingly recognized that, without the contribution of evolutionary biology, ecological predictions of the effects of global change can be naive (Travis and Futuyma 1993). The information base and the expertise currently exist to apply evolutionary knowledge to global change concerns. With this paper, we hope to stimulate efforts in that direction.
Responses to this article are invited. If accepted for publication, your response will be hyperlinked to the article. To submit a comment, follow this link. To read comments already accepted, follow this link.
Acknowledgments
We are indebted to Mauro Santos for insightful suggestions, and we thank Judy Bronstein and several anonymous reviewers for their comments on earlier drafts of this paper. Both F. Rodríguez-Trelles and M. A. Rodríguez were supported by the Spanish Ministry for Education and Culture (Contratos de Reincorporación). Additional support was provided for F. Rodríguez-Trelles by the Comunidad Autónoma de Madrid (grant: COR-0077/94 to A. Gómez-Sal), and for M. A. Rodríguez by the Spanish CICYT (grant: AMB96-1161 to A. Gómez-Sal).
Aquadro, C. F., A. L. Weaver, S. W. Schaeffer, and W. W. Anderson. 1991. Molecular evolution of inversions in Drosophila pseudoobscura: the amylase gene region. Proceedings of the National Academy of Sciences (USA) 88:305-309.
Bella, D. A., R. Jacobs, and L. Hiram. 1994. Ecological indicators of global climate change: a research framework. Environmental Management 18:489-500.
Brncic, D. 1994. The colonization of Chile by Drosophila subobscura and its consequences. Pages 154-169 in L. Levine, editor. Genetics of natural populations: the continuing importance of Theodosius Dobzhansky. Columbia University Press, New York, New York, USA.
Crow, J. F., and M. Kimura. 1970. An introduction to population genetics theory. Harper and Row, New York, New York, USA.
Davis, A. J., L. S. Jenkinson, J. H. Lawton, B. Shorrocks, and S. Wood. 1998. Making mistakes when predicting shifts in species range response to global warming. Nature 391:783-786.
Davis, M. B. 1981. Quaternary history and the stability of forest communities. Pages 132-153 in D. C. West, H. H. Shugart, and D. B. Botkin, editors. Forest succession. Springer-Verlag, New York, New York, USA.
Dobzhansky, T. H. 1947. A response of certain gene arrangements in the third chromosome of Drosophila pseudoobscura to natural selection. Genetics 32:142-160.
Endler, J. A. 1986. Natural selection in the wild. Princeton University Press, Princeton, New Jersey, USA.
Feder, M. E., N. V. Cartano, L. Milos, R. A. Krebs, and S. L. Lindquist. 1996. Effect of engineering Hsp70 copy number of Hsp70 expression and tolerance of ecologically relevant heat shock in larvae and pupae of Drosophila melanogaster. Journal of Experimental Biology 199:1837-1844.
Hilbish T. J., and R. K. Koehn. 1985. The physiological basis of natural selection at the LAP locus. Evolution 39:1302-1317.
Holt, R. D. 1990. The microevolutionary consequences of climate change. Trends in Ecology and Evolution 5:311-315.
_______ . 1995. Linking species and ecosystems: where's Darwin? Pages 273-279 in C. G Jones and J. H Lawton, editors. Linking species and ecosystems. Chapman and Hall, New York, New York, USA.
Jenkinson, L. S., A. J. Davis, S. Wood, B. Shorrocks, and J. H. Lawton. 1996. Not that simple: global warming and predictions of insect ranges and abundances - results from a model insect assemblage in replicated laboratory ecosystems. Aspects of Applied Biology 45:343-348.
Kingsolver, J. G., R. B. Huey, and P. M. Kareiva. 1993. An agenda for population and community research on global change. Pages 480-486 in P. M. Kareiva, J. G. Kingsolver, and R. B. Huey, editors. Biotic interactions and global change. Sinauer Associates, Sunderland, Massachusetts, USA.
Kingsolver, J. G., and W. B. Watt. 1983. Thermoregulatory strategies in Colias butterflies: thermal stress and the limits to adaptation in temporally varying environments. American Naturalist 121:32-55.
Krimbas, C. B. 1992. The inversion polymorphism of Drosophila subobscura. Pages 127-220 in C. B. Krimbas and J. R. Powell, editors.Drosophila inversion polymorphism. CRC Press, Boca Raton, Florida, USA.
Krimbas, C. B., and M. Loukas. 1980. The inversion polymorphism of Drosophila subobscura. Evolutionary Biology 12:163-234.
Krimbas, C. B., and J. R. Powell, editors. 1992. Drosophila inversion polymorphism. CRC Press, Boca Raton, Florida, USA.
Lande, R., and S. Shannon. 1996. The role of genetic variation in adaptation and population persistence in a changing environment. Evolution 50:434-437.
Lynch, M., J. Conery, and R. Bürger. 1995. Mutation accumulation and the extinction of small populations. American Naturalist 146:489-518.
Mennozzi, P., and C. B. Krimbas. 1992. The inversion polymorphism of Drosophila subobscura revisited: synthetic maps of gene arrangements frequencies and their interpretation. Journal of Evolutionary Biololgy 5:625-641.
Misra, R. K., and E. C. R. Reeve. 1964. Clines in body dimensions in populations of Drosophila subobscura. Genetical Research 5:240-254.
Moltó, M. D., L. Pascual, M. J. Martínez-Sebastián, and R. de Frutos. 1992. Genetic analysis of heat shock response in the three Drosophila species of the obscura group. Genome 35:870-880.
Parmesan, C. 1997. Climate and species' range. Nature 382:765-766.
Parsons, P. A. 1991. Biodiversity conservation under global climate change: the insect Drosophila as a biological indicator? Global Ecology and Biogeography Letters 1:77-83.
Peters, R. H., and R. de Bernardi. 1987. Daphnia. Instituto Italiano di Idrobiologia, Pallanza, Italy.
Prevosti, A. 1966. Chromosomal polymorphisms in Western Mediterranean populations of Drosophila subobscura. Genetical Research 7:149-158.
Prevosti, A., G. Ribó, L. Serra, M. Aguadé, J. Balaña, M. Monclús, and F. Mestres. 1988. Colonization of America by Drosophila subobscura: experiment in natural populations that supports the adaptive role of chromosomal-inversion polymorphism. Proceedings of the National Academy of Sciences (USA) 85:5597-5600.
Richardson, A. M. M. 1974. Differential climatic selection in natural populations of land snail Cepaea nemoralis. Nature 247:572-573.
Rodríguez-Trelles, F., G. Álvarez, and C. Zapata. 1996. Time-series analysis of seasonal changes of the O inversion polymorphism of Drosophila subobscura. Genetics 142:179-187.
Rodríguez-Trelles, F., and M. A. Rodríguez. 1998. Rapid microevolution and loss of chromosomal diversity in Drosophila in response to climate-warming. Evolutionary Ecology 12(7), in press.
Saura, A. 1994. Genetic load and population size in northern populations of Drosophila subobscura. Pages 173-187 in L. Levine, editor. Genetics of natural populations: the continuing importance of Theodosius Dobzhansky. Columbia University Press, New York, New York, USA.
Sawyer, L. A., J. M. Hennessy, A. A. Peixoto, E. Rosato, H. P. Parkinson, R. Costa, and C. P. Kyriacou. 1997. Natural variation in a Drosophila clock gene and temperature compensation. Science 278:2117-2120.
Schneider, S. H. 1993. Scenarios of global-warming. Pages 234-250 in P. M. Kareiva, J. G. Kingsolver, and R. B. Huey, editors. Biotic interactions and global change. Sinauer Associates, Sunderland, Massachusetts, USA.
Schowalter, T. D., W. W. Hargrove, and D. A. Crossley, Jr. 1986. Herbivory in forested ecosystems. Annual Review of Entomology 31:177-196.
Sturtevant, A. H., and Th. Dobzhansky. 1936. Inversions in the third chromosome of wild races of Drosophila pseudoobscura, and their use in the study of the history of the species. Proceedings of the National Academy of Sciences (USA) 22:448-450.
Templeton, A. R., H. Hollocher, S. Lawyer, and J. S. Johnston. 1990. The ecological genetics of abnormal abdomen in Drosophila mercatorum. Pages 17-35 in J. S. F. Barker, W. T. Starmer, and R. J. MacIntyre, editors. Ecological and evolutionary genetics of Drosophila. Plenum Press, New York, New York, USA.
Travis, J., and D. J. Futuyma. 1993. Global change: lessons from and for evolutionary biology. Pages 251-263 in P. M. Kareiva, J. G. Kingsolver, and R. B. Huey, editors. Biotic interactions and global change. Sinauer Associates, Sunderland, Massachusetts, USA.
Vitousek, P. M. 1992. Global environmental change: an introduction. Annual Review of Ecology and Systematics 23:1-14.
Watt, W. B., R. C. Cassin, and M. S. Swam. 1983. Adaptation at specific loci. III. Field behavior and survivorship differences among Colias PGI genotypes are predictable from in vitro biochemistry. Genetics 103:725-739.
White, M. J. D. 1978. Modes of speciation. W. H. Freeman, San Francisco, California, USA.
Wright, S., and T. H. Dobzhansky. 1946. Experimental reproduction of some of the changes caused by natural selection in certain populations of Drosophila pseudoobscura. Genetics 31:125-156.
List of the studies containing frequency data of Drosophila subobscura chromosomal inversion polymorphisms in natural populations throughout the Palearctic.
1. Andjelkovic, M., and D. Sperlich. 1973. Inversion polymorphism in a Pannonian population of Drosophila subobscura. Egyptian Journal of Genetics and Cytology 2: 144-147.
2. Bächli, G., H. Burla, and H. Jungen. 1982. Drosophila subobscura in Egypt and its probable derivation. Genetica 59: 3-7.
3. Brehm, A., and C. B. Krimbas. 1988. The inversion polymorphism of Drosophila subobscura natural populations from Portugal. Genética Ibérica 39: 235-248.
4. Burla, H., and W. Götz. 1965. Veränderlichkeit des chromosomalen Polymorphismus bei Drosophila subobscura. Genetica 36: 83-104.
5. Burla, H., H. Jungen, and G. Bächli. 1986. Population structure of Drosophila subobscura: nonrandom microdispersion of inversion polymorphism on a mountain slope. Genetica 70: 9-15.
6. Charlesworth, B., D. Charlesworth, and M. Loukas. 1979. A study of linkage disequilibrium in British populations of Drosophila subobscura, with an appendix by K. Morgan. Genetics 92: 983-994.
7. de Frutos, R. 1972. Contribution to the study of chromosomal polymorphism in the Spanish populations of Drosophila subobscura. Genética Ibérica 24: 123-140.
8. de Frutos, R., and A. Prevosti. 1984. Temporal changes of chromosomal polymorphism in natural populations of Drosophila subobscura. Genetica 63: 181-187.
9. Fontdevila, A., C. Zapata, G. Álvarez, L. Sánchez, J. Méndez, and I. Enríquez. 1983. Genetic coadaptation in the chromosomal polymorphism of Drosophila subobscura. I. Seasonal changes of gametic disequilibrium in a natural population. Genetics 105: 935-955.
10. García, M. P., and A. Prevosti. 1981. Association between allozyme alleles and chromosomal arrangements of the O chromosome in Drosophila subobscura. I. Data of natural populations. Genética Ibérica 33: 151-174.
11. Goldschmidt, E. 1956. Chromosomal polymorphism in a population of Drosophila subobscura from Israel. Journal of Genetics 54: 474-496.
12. Goldschmidt, E. 1956. Structural polymorphism in the Israel race of Drosophila subobscura. Bulletin of the Research Council of Israel 5B(3-4): 316-317.
13. Goldschmidt, E. 1958. Polymorphism and coadaptation in natural populations of Drosophila subobscura. Proceedings of the 10th International Congress of Entomology (Montreal) 2: 821-828.
14. Gosteli, M. 1990. Chromosomal polymorphism in natural populations of Drosophila subobscura near Zurich, Switzerland: a contribution to long-term comparisons. Genetica 81: 199-204.
15. Gosteli, M. 1991. Differential flight activity among karyotypes: daily and weather-induced changes in chromosomal inversion polymorphism in natural populations of Drosophila subobscura. Genetica 84: 129-136.
16. Götz, W. 1965. Chromosomaler polymorphismus in einem muster von Drosophila subobscura aus Marokko, mit darstellung der heterozygotieverhältnisse als heterozygotiediagram. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 97: 40-45.
17. Götz, W. 1967. Untersuchungen über den chromosomalen Strukturpolymorphismus in kleinasiatischen und persischen Populationen von Drosophila subobscura Collin. Molecular and General Genetics 100: 1-38.
18. Jungen, H. 1966. Chromosomal polymorphism in a natural population of Drosophila subobscura from Tunis. Drosophila Information Service 41: 173.
19. Jungen, H. 1968. Inversions polymorphismus in tunesischen Populationen von Drosophila subobscura Collin. Archiv. Julius Klaus-stiftung für Vererbungsforschung, Sozialanthropolgie und Rassen 43: 1-55.
20. Knight, G. R. 1961. Structural polymorphism in Drosophila subobscura Coll. from various localities in Scotland. Genetical Research 2: 1-9.
21. Krimbas, C. B. 1992. The inversion polymorphism of Drosophila subobscura. Pages 127-220 in C. B. Krimbas and J. R. Powell, editors. Drosophila inversion polymorphism. CRC Press. Boca Raton, Florida, USA.
22. Krimbas, C. B. 1963. Inversion polymorphism in natural populations of Drosophila subobscura in Greece. Drosophila Information Service 37: 95-96.
23. Krimbas, C. B. 1964. The genetics of Drosophila subobscura populations. II. Inversion polymorphism in a population from Holland. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 95: 125-128.
24. Krimbas, C. B. 1965. Further data on inversion polymorphism of Drosophila subobscura in Greece. Drosophila Information Service 40: 45.
25. Krimbas, C. B. 1965. The genetics of Drosophila subobscura populations. I. Inversion polymorphism in populations of southern Greece. Evolution 18: 541-552.
26. Krimbas, C. B. 1967. The genetics of Drosophila subobscura populations. III. Inversion polymorphism and climatic factors. Molecular and General Genetics 99: 133-150.
27. Krimbas, C. B. 1971. Gene arrangement frequencies in Pindos population of D. subobscura. Drosophila Information Service 47: 84.
28. Krimbas, C. B., and J. R. Powell, editors. 1992. Drosophila inversion polymorphism. CRC Press. Boca Raton, Florida, USA.
29. Krimbas, C. B., and V. Alevizos. 1973. Appendix in A. Saura's, S., J. Lakovaara, J. Lokki, and P. Lankinen. Genic variation in central and marginal populations of Drosophila subobscura. Hereditas 75: 44.
30. Krimbas, C. B., and V. Alevizos. 1973. The genetics of Drosophila subobscura populations. IV. Further data on inversion polymorphism in Greece. Evidence for microdifferentiation. Egyptian Journal of Genetics and Cytology 2: 121-132.
31. Krimbas, C. B., and M. Loukas.1979. Inferred gene arrangements in 14 natural populations of Drosophila subobscura. European Drosophila Population Biology Group Bulletin 3: 1-7.
32. Krimbas, C. B., and M. Loukas. 1980. The inversion polymorphism of Drosophila subobscura. Evolutionary Biology 12: 163-234.
33. Krimbas, C. B., M. Loukas, and E. Diamantopoulou. 1971. Gene arrangement and gene frequencies in Mt. Parnes population of Drosophila subobscura. Drosophila Information Service 46.
34. Krimbas, C.B. 1963. Inversion polymorphism in southern Greek populations of Drosophila subobscura. Drosophila Information Service 38: 81.
35. Kunze-Mühl, E., E. Müller, and D. Sperlich. 1958. Qualitative, quantitative und jahreszeitliche untersuchungen über den chromosomalen polymorphismus natürlicher populationen von Drosophila subobscura Coll. in der umgebung vo wien. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 89: 636-646.
36. Kunze-Mühl, E., and E. Müller. 1958. Weitere Untersuchungen über die chromosomale Struktur und natürlichen Strukturtypen von Drosophila subobscura. Chromosoma (Berlin) 9: 559-570.
37. Kunze-Mühl, E., and D. Sperlich. 1955. Inversionen und chromosomale Strukturtypen bei Drosophila subobscura Coll. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 87: 65-84.
38. Kunze-Mühl, E., and D. Sperlich. 1962. Vergleichende Untersuchungen über den chromosomalen Strukturpolymorphismus in Insel- und Festland-Populationen von Drosophila subobscura. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 93: 237-248.
39. Larruga, J. M., V. M. Cabrera, A. M. González, and A. Gullón. 1983. Molecular and chromosomal polymorphism in continental and insular populations from the south-western range of D. subobscura. Genetica 60: 191-205.
40. Larruga, J. M., J. Rozas, M. Hernández, A. M. González, and V. M. Cabrera. 1993. Latitudinal differences in sex chromosome inversions, sex- linked allozymes, and mitochondrial DNA variation in Drosophila subobscura. Genetica 92: 67-74.
41. Loukas, M., and C. B. Krimbas. 1975. The genetics of Drosophila subobscura populations. V. A study of linkage disequilibrium in natural populations between genes and inversions of the E chromosome. Genetics 80: 331-347.
42. Loukas, M., C. B. Krimbas, and K. Morgan. 1980. The genetics of Drosophila subobscura populations: XVI. Further data on linkage disequilibrium. Genetics 95: 757-768.
43. Loukas, M., C. B. Krimbas, and Y. Vergini. 1979. The genetics of Drosophila subobscura populations. IX. Studies on linkage disequilibrium in four natural populations. Genetics 93: 497-523.
44. Loukas, M., Y. Vergini, and C. B. Krimbas. 1981. The genetics of Drosophila subobscura populations. XVII. Further genetic heterogeneity within electromorphs by urea denaturation. The effect of increased genetic variability on linkage disequilibrium studies. Genetics 97: 429-441.
45. Malogolowkin-Cohen, Ch. 1980. Inversion polymorphism in Drosophila subobscura in Israel. Drosophila Information Service 55: 90.
46. Malogolowkin-Cohen, Ch., and D. Sperlich. 1981. The effect of isolation and marginality on the inversion polymorphism of Drosophila subobscura in Israel. Revista Brasileira de Genetica 4: 213-230.
47. Martínez-Sebastián, M. J., A. Lattore, and R. de Frutos. 1984. Chromosomal polymorphismn of "Drosophila subobscura" populations sampled at different altitudes. Genética Ibérica 36: 1-12.
48. Mennozzi, P., and C. B. Krimbas. 1992. The inversion polymorphism of Drosophila subobscura revisited : synthetic maps of gene arrangements frequencies and their interpretation. Journal of Evolutionary Biology 5: 625-641.
49. Mestres, F., J. Balanya, C. Segarra, and L. Serra. 1989. Study of the association between the O5 chromosomal inversion and a lethal gene in paleartic and colonizing populations of D. subobscura. Abstracts 11th European Drosophila Research Conference, Marseilles, 45.
50. Mestres, F., J. Balanya, C. Segarra, A. Prevosti, and L. Serra. 1994. O chromosome inversion polymorphism in Northern and Atlantic Europe and its implications in the American colonization by Drosophila subobscura. Zeitschrift fuer Zoologische Systematik und Evolutionsforschung 32: 108-116.
51. Orengo, D-J., and A. Prevosti. 1996. Temporal changes in chromosomal polymorphisms of Drosophila subobscura related to climate changes. Evolution 50: 1346-1350.
52. Pentzos-Daponte, A. 1964. Qualitative und quantitative Untersuchungen über den chromosomalen Polymorphismus natürlicher Populationen von Drosophila subobscura in der Umgebung von Thessaloniki (Griechenland). Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 95: 129-144.
53. Pentzos-Daponte, A. 1964. Qualitative und quantitative untersuchungenüber den chromosomalen polymorphismus naturlicher populationen von Drosophila subobscura. Zeitschrift für Vererbungslehre 97: 46-54.
54. Pinsker, W., and D. Sperlich. 1979. Allozyme variation in natural populations of Drosophila subobscura along a north-south gradient. Genetica 50: 207-214.
55. Pinsker, W., P. Lankinen, and D. Sperlich. 1978. Allozyme and inversion polymorphism in a central population of Drosophila subobscura. Genetica 48: 207-219.
56. Prevosti, A. 1966. Chromosomal polymorphisms in Western Mediterranean populations of Drosophila subobscura. Genetical Research 7: 149-158.
57. Prevosti, A. 1964. Tipos cromosómicos de Drosophila subobscura en una población de Lagrasse (Francia). Genética Ibérica 16: 1-19.
58. Prevosti, A. 1964. Chromosomal polymorphism in Drosophila subobscura populations from Barcelona (Spain). Genetical Research 5: 27-38.
59. Prevosti, A. 1966. Chromosomal polymorphism in Western Mediterranean populations of Drosophila subobscura. Genetical Research 7: 149-158.
60. Prevosti, A. 1968. Efecto de la cordillera pirenaica sobre la distribución geográfica de las ordenaciones cromosómicas de Drosophila subobscura. Pirineos 79-80: 221-228.
61. Prevosti, A. 1971. Chromosomal polymorphism in Drosophila subobscura Coll. populations from the Canary Islands. Genética Ibérica 23: 69-84.
62. Prevosti, A. 1972. Chromosomal polymorphism in Drosophila subobscura populations from the Madeira Island. Genética Ibérica 24: 11-21.
63. Prevosti, A. 1974. Chromosomal inversion polymorphism in the southwestern range of Drosophila subobscura distribution area. Genetica 45: 111-124.
64. Prevosti, A., R. de Frutos, G. Alonso, A. Latorre, M. Monclús, and M. J. Martínez. 1984. Genetic differentiation between natural populations of Drosophila subobscura in the Western Mediterranean area with respect to chromosomal variation. Genetics, Selection and Evolution 16: 143-156.
65. Rodríguez-Trelles, F., G. Álvarez, and C. Zapata. 1996. Time-series analysis of seasonal changes of the O inversion polymorphism of Drosophila subobscura. Genetics 142: 179-187.
66. Sperlich, D. 1961. Untersuchungen über den chromosomalen polymorphismus einer population von Drosophila subobscura auf den Liparischen Inseln. Zeitschrift für Vererbungslehere 92: 74-84.
67. Sperlich, D. 1964. Chromosomale Strukturanalysen und Fertilitätsprüfung an einer marginal Population von Drosophila subobscura. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 95: 73-81.
68. Sperlich, D., and E. Kunze-Mühl. 1963. Der chromosomale polymorphismus einer population von Drosophila subobscura auf der Insel Ustica mi Vergleich mit anderen Insel- und Festland-standorten. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 94: 94-100.
69. Sperlich, D., and H. Feuerbach. 1966. Ist der chromosomale Struktur-polymorphismus von Drosophila subobscura stabil oder flexibel? Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 98: 16-24.
70. Sperlich, D., W. Pinsker, and V. G. Mitrofanov. 1981. Genetic characterization of a natural population of Drosophila subobscura from the Northern Caucasus (USSR) in comparison with other population samples. Genetica 54: 329-334.
71. Stumm-Zollinger, E. 1953. Vergleichende Untersuchung über die Inversionshaufigkeit bei Drosophila subobscura in Populationen der Schweiz und Sudwesteuropas. Zeitchsrift Induktive Abstammungs-u. Vererbungslehre 85: 382-407.
72. Stumm-Zollinger, E., and E. Goldschmidt. 1959. Geographical differentiation on inversion systems in Drosophila subobscura. Evolution 13: 89-98.
73. Zivanovic, G., M. Milanovic, and M. Andjelkovic. 1995. Chromosomal inversion polymorphism of Drosophila subobscura populations from Jastrebc mountain shows temporal and habitat-related changes. Journal of Zoological Systematics and Evolutionary Research 33: 81-83.
74. Zivanovic, G., M. Milanovic, and M. Andjelkovic. 1995. Inversion polymorphism in Drosophila subobscura from Jastrebac Mountain and localization of the gene arrangement O3+4+1 distribution. Drosophila Information Service 76: 91-92.
75. Zouros, E., C. B. Krimbas, S. Tsakas, and M. Loukas. 1974. Genic versus chromosomal variation in natural populations of Drosophila subobscura. Genetics 78:1223-1244. (with an appendix by M. Loukas, S. Tsakas, C. B. Krimbas, E. Zouros, E. Diamantopoulou-Panopoulou, and V. Alevizos).
Address of Correspondent:
Francisco Rodríguez-Trelles
Departament de Genètica i de Microbiología,
Universitat Autonoma de Barcelona
Edifici C
08193 - Bellaterra
Barcelona, Spain
Phone: 34-93-581-2725
Fax: 34-93-581-2387
ibge2@blues.uab.es
*The copyright to this article passed from the Ecological Society of America to the Resilience Alliance on 1 January 2000.